The coverage of Bin79 and sequence identity between the mapped reads and genome sequence were determined. 17 May, 2017) with the default parameters and idtag option. The reads from 2Pm were aligned to the Cellvibrio sp. The manually checked sequences were blasted against the NCBI NT database with the cutoff E-value 1e−5, and the scaffolds with no Cellvibrio hits were discarded. If a sequence region was not well supported by the mapped paired-end reads (i.e., the paired-end reads were not concordantly mapped or the average coverage was below five-fold in that region or the coverage in the region was significantly different from the average coverage of the genome), the affected region was discarded. The metagenomic reads from sample 2Pm (rhizoplane of tree 2, the sample with the highest abundance of Bin79) were mapped back to the refined scaffolds using Bowtie2, and the alignments were carefully checked using the CLC Genomics Workbench ver. 
The contigs of Bin79 were scaffolded using the Medusa server based on the five publicly available Cellvibrio genomes (accessed 2 January, 2017), including C. Of note, we observed that Cellvibrio exhibited significantly higher relative abundance in the citrus rhizoplane than in the rhizosphere. In our previous study, we surveyed the citrus rhizosphere microbiome at the global scale and observed that Cellvibrio was always enriched in the citrus rhizosphere compared with the corresponding bulk soil, regardless of citrus genotype, soil type, and climate, probably resulting from the more abundant plant-driven polysaccharides in the rhizosphere compared with those of the bulk soil. Under such selective forces, only certain microbes from the rhizosphere microbiome can be further enriched and colonize the rhizoplane niche, leading to lower diversity and density of the microbial community in the rhizoplane compared with that of the rhizosphere. Plants have developed active cell wall integrity-monitoring systems to sense cell wall damage-associated signals, and complex defense responses, including production of reactive oxygen species and activation of hormone-mediated immune responses, will be triggered once the signals are detected. Degradation of the plant cell wall is an important strategy for pathogens to infect the plant host. Plant cells are surrounded by a thick cell wall primarily composed of complex carbohydrates, including cellulose, hemicellulose, lignin, and pectin, and the cell wall plays critical roles in protecting plants against biotic and abiotic stresses, such as pathogen attack and drought. The influences imposed by the plant host on the rhizosphere microbiome are relatively low however, the root surface, i.e., the rhizoplane, poses different challenges for microorganisms to colonize from the rhizosphere. Mainly owing to root excretion and sloughing, gradients of carbon sources, phytochemicals, as well as other nutrients are formed from the root surface to the surrounding soil, and certain microorganisms are attracted from bulk soil towards roots and form the rhizosphere microbiome. Overall, this study characterizes a novel Cellvibrio strain and indicates the mechanisms involved in its adaptation to the rhizoplane from meta-omics data without cultivation. Comparative genomics analyses with five related Cellvibrio strains showed the importance of gene gain events for the rhizoplane adaptation of Bin79. 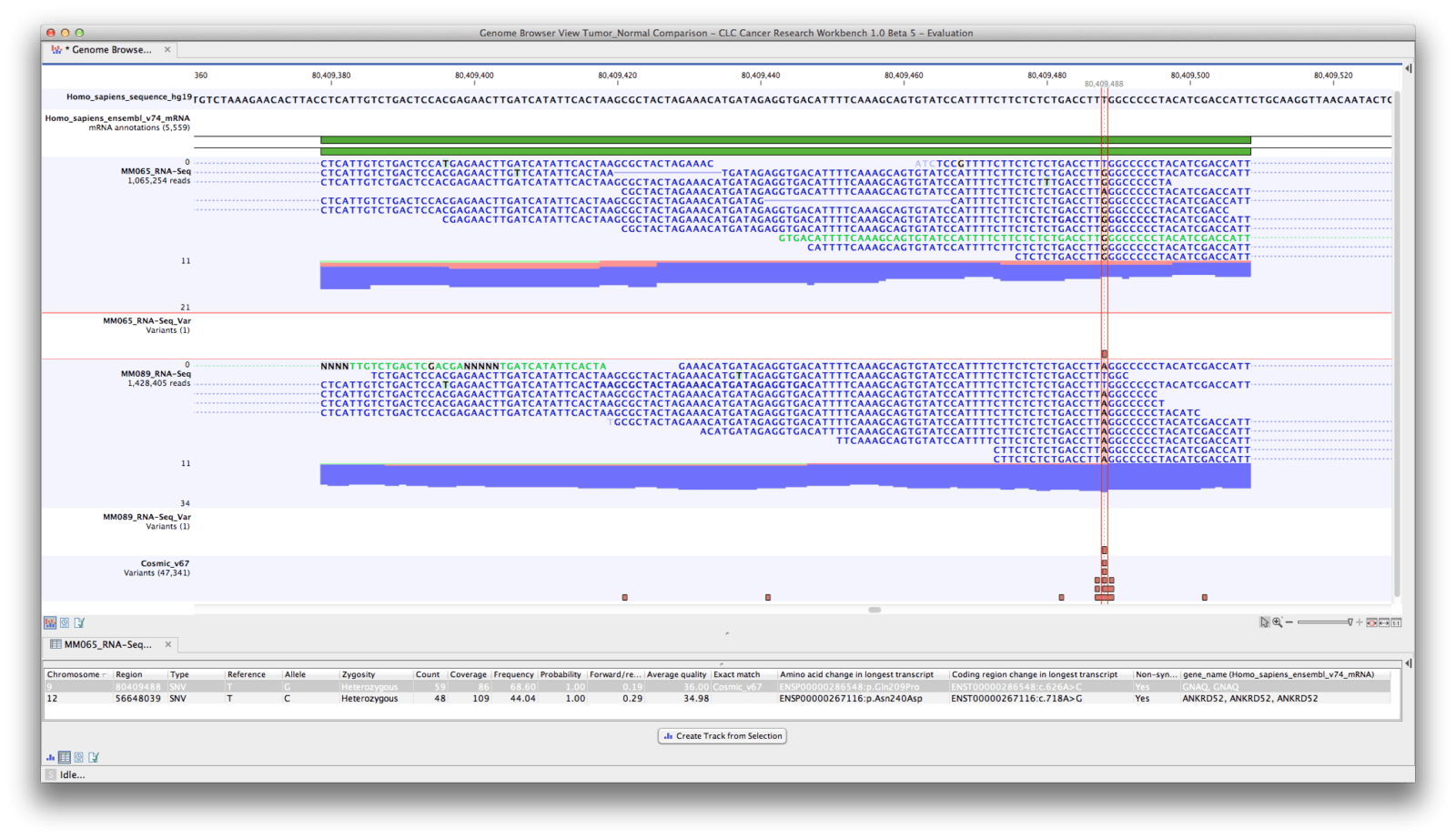
In addition, genes involved in repelling plant immune responses were significantly activated in the rhizoplane. Enhanced expression of multi-drug efflux genes and iron acquisition- and storage-associated genes in the rhizoplane indicated mechanisms by which Bin79 competes with other microbes. Differential gene expression analysis demonstrated that plant cell wall degradation genes were repressed, whereas genes encoding chitin-degrading enzymes were induced in the rhizoplane compared with the rhizosphere. We recovered a near-complete metagenome-assembled genome representing a potentially novel species of Cellvibrio, herein designated Bin79, with genome size of 5.71 Mb across 11 scaffolds. Here we investigated the mechanisms underlying the rhizosphere-to-rhizoplane enrichment of Cellvibrio through genome-centric metagenomics and metatranscriptomics analyses. Maintaining integrity of the plant cell walls is critical for plant health, however, our previous study showed that Cellvibrio, which is recognized by its robust ability to degrade plant cell walls, was enriched from the citrus rhizosphere to the rhizoplane (i.e., the root surface).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
